|




|
The purpose of our program is to provide the optimal set of mixing matrix and starting sequences in order to maximize
the yield of specific target motifs in in vitro selection. You may choose the target RNA structure for
in vitro selection. Our program will then give you the optimal set of mixing matrix and starting sequences.
See the tutorial section for background details.
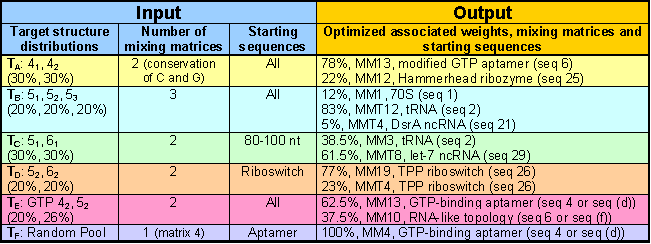
As detailed above, the RNA pool designer computes the optimal designed pool parameters corresponding to the user input. For example, if the user requests to use two mixing matrices with the conservation of C and G and all starting sequences to achieve 30% each of 41 and 42 tree motifs as in target Pool TA, the optimization specifies 78% of mixing matrix 13 with modified GTP aptamers (seq 5) as starting sequence and 22% of mixing matrix 12 with the hammerhead ribozyme (seq 25). This combination yields the desired structural distribution (subject to the correction of the 2D structure data).
Similarly, users can construct target Pool TB to use three mixing matrices and all starting sequences to find 20% each of 51, 52 and 53 structures. The optimized output specifies 12% of mixing matrix 1 with 70S (seq 1) as starting sequence, 83% of MMT 12 with tRNA (seq 2), and 5% of MMT 4 with DsrA ncRNA (seq 21).
Users can further filter the output by selecting additional options to choose certain groups of starting sequences. In target Pool TC, the user requests to use two mixing matrices to achieve 30% each of 51 and 61 similar to target Pool TA. However, the user is limiting the results by choosing starting sequences with 80-100 nucleotides. The optimized output shows 38.5% of mixing matrix 3 with tRNA (seq 2) and 61.5% of MMT 8 with let-7 ncRNA (seq 29). Similarly, target Pool TD further filters the optimized results by limiting the possible starting sequences to have riboswitch function.
We apply our pool design approach for enhancing GTP-binding aptamers. Szostak’s group recently found that the GTP aptamer’s
binding affinity is correlated with structural complexity (e.g., number of stems) (Carothers et al., 2004;Carothers et al., 2006).
Specifically, a high affinity GTP aptamer (panel C in Ref. (Carothers et al., 2004)) is related to the
52 tree structure. Interestingly, no GTP aptamer with a
42 tree structure has been reported although it is structurally similar to the
52 tree. Because the frequency of the 42 motif is only 12% in random pool TF, we propose designing a
GTP aptamer pool by enriching the pool with 52 and 42 motifs. Our target pool fractions (Ti) are
20% for 42 and 26% for 52.
Our optimization yields Pool TE as a combination of two subpools:
(sequence (d), MM13, 0.625) and
(sequence (f), MM10, 0.375).
Experimental synthesis of designed pools can be performed by using those sets. The frequencies of 42 and 52 trees
in the designed pool are 21% and 26% respectively, nearly as desired and very different for 12% and 16% distributions of these motifs
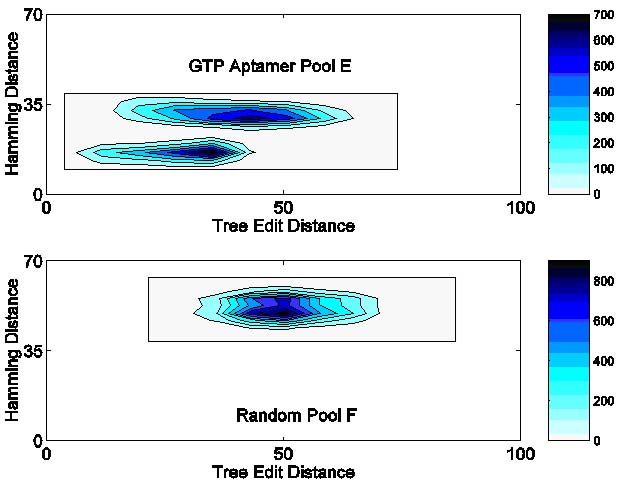
in random pool TF. The following sequence/structure contour plots show differences between the designed and random pools;
the designed pool has a relatively high mean mutation rate of 0.349.