



|




|
|
In vitro selection is an experimental approach that allows the screening of large (~1015) random-sequence
libraries of nucleic acid molecules for a specific function, such as binding or catalysis. In recent years, numerous
target-binding RNA molecules (aptamers) and novel RNA enzymes
(ribozymes) have been identified by in vitro selection.
The typical aptamer selection scheme is provided as follows:
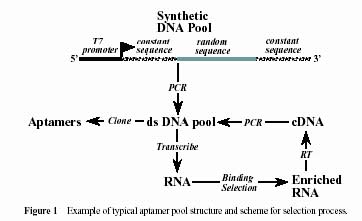 Figure taken from Wilson DW and Szostak JW. Ann. Rev. Biochem., 68:611-648 (1999) The standard experimental protocol involves synthesizing DNA sequences and then transcribing them into RNA sequences by RNA polymerases. In random pools, the length of sequence is fixed and each position has an equal probability of A, U, C and G. Computer simulations of this process show that RNA topology distributions in random pools with different lengths from 25nt to 100nt are not uniform. For example, simple linear tree structures are abundant, whereas complex structures are virtually absent in random pools. | |
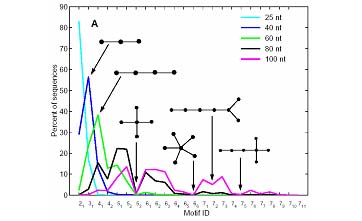 Figure 2. RNA topology distributions in random pools. Figure taken from Gevertz J, Gan HH, and Schlick T. RNA 11:853 (2005)
| |
