



|




|
|
The pool structural distribution is calculated by mapping RNA secondary structures into graph space. This is done by
predicting secondary structures of all sequences using the Vienna RNA fold package and then converting them into
tree graphs. Specifically, base pairing information in the .ct file
generated by the RNAfold program is used to convert a secondary fold into a tree graph. The topologies of the folds
are determined using Laplacian eigenvalues of tree graphs as implemented
in our RNA Matrix Program.
|
Pool Sizes10,000 sequence pools are adequate for assessing structural distributions using simple tree graphs. To show the effect of pool size, we compute the frequency of several tree motifs (41, 42, 51, 52, 53, and 61) for pool sizes of 5,000-60,000 sequences using MM4 and the initial tRNA sequence.
The pool fractions for distinct tree motifs saturate beyond 5,000 sequences, indicating that the error due to sample size is small. Structure prediction and conversion to tree graphs for 10,000 80-nt sequences require about 1 hour on SGI 300 MHz MIPS R12000 IP27 processor.
|
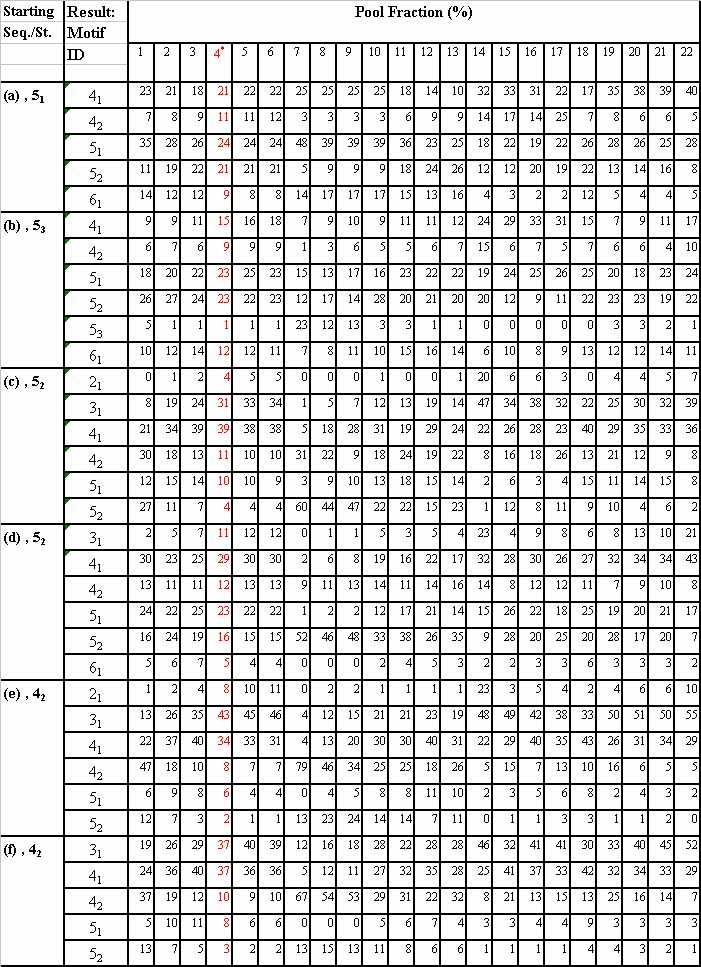
ExamplesStructural distributions of pools generated by 22 mixing matrices starting with six sequences (a)-(f). Yellow shaded numbers represent frequencies greater than those in the random pool (MM4).
 |
